New York Variant --The E484K or S477N Mutations
B.1.526 contains either the E484K or the S477N mutation
"...Caltech researchers discovered the rise in B.1.526 by scanning for mutations in hundreds of thousands of viral genetic sequences in a database called GISAID. “There was a pattern that was recurring, and a group of isolates concentrated in the New York region that I hadn’t seen,” said Anthony West, a computational biologist at Caltech. He and his colleagues found two versions of the coronavirus increasing in frequency: one with the E484K mutation seen in South Africa and Brazil, which is thought to help the virus partially dodge the vaccines; and another with a mutation called S477N, which may affect how tightly the virus binds to human cells..." NY times Feb 24, 2021
SARS-CoV-2 lineage B.1.526 emerging in the New York region detected by software utility created to query the spike mutationallandscape West el al., Caltech
The most commonsetsof spike mutationsin this lineage (now designated as B.1.526) areL5F,T95I,D253G,E484Kor S477N,D614G, andA701V.This lineage appeared in late November 2020, and isolates from this lineage account for ~25% of coronavirus genomes sequenced and deposited from New York during February 2021
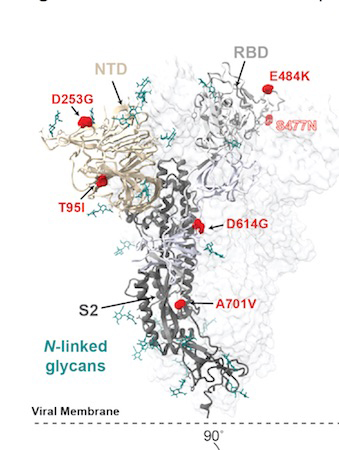
Image above: West et al., Feb 23, 2021
Covid-19: The E484K mutation and the risks it poses
The E484K mutation occurs in the receptor-binding domain (RBD) that the virus uses to bind to the human ACE2 receptor and has been associated with escape from neutralizing antibodies.
There has been research showing that the current vaccines work against the UK B.1.1.7 variant without the E484K mutation. However, recent clinical trials by Novavax and Johnson & Johnson showed that their new vaccines were less effective in South Africa compared with the UK or US, which is presumably because of the high level of virus carrying the E484K mutation.
"...E484 is the site in the RBD where mutations usually have the largest effect on binding andneutralization..." source
See: B.1.351, South Africa (E484K variant for more information on the E484K mutation and how it affects conformation of spike protein.
The E484K variants and Vaccines
The E484K mutation in the SARS-CoV-2 spike protein reduces but does not abolish neutralizing activity of human convalescent and post-vaccination sera -- "...human sera with high neutralization titers against the USA-WA1/2020 strain were still able to neutralize the E484K rSARS-CoV-2. Therefore, it is important to aim for the highest titers possible induced by vaccination to enhance protection against newly emerging SARS-CoV-2 variants. Two vaccine doses may be needed for induction of high antibody titers against SARS-CoV-2. ..."
"...Based on the computational analysis of mutational effects, the D614G, N501Y and S477N mutations decreased stability and were tolerated. For disease propensity prediction, N501Y was more prone to disease compared to D614G, while S477N was not prone to disease. The mutation of D to G at position 614 and S to N at position 477 for secondary structure prediction shows no changes in secondary structure while remaining in the coil region, whereas the mutation of N to Y at position 501 changes from coil structure to extended strand. N501Y mutation has a higher affinity to human ACE2 protein compared to D614G and S477N based on a docking study...."
S477N has less bonding to others and reduces stability...."
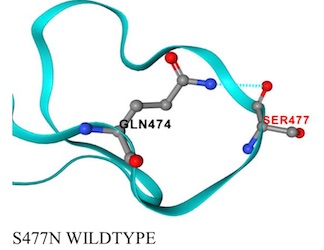
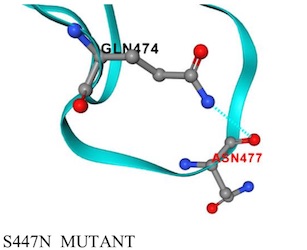
-------------->spin on -------->- spin off
To Rotate the Molecule--->Left Click and Drag
To Zoom-->>Left Click + hold Shift button and Drag Vertically
Jmol Menu --->>Right-Click
-------------->spin on -------->- spin off
To Rotate the Molecule--->Left Click and Drag
To Zoom-->>Left Click + hold Shift button and Drag Vertically
Jmol Menu --->>Right-Click
For complete details of the E484K mutation see the South African variant page
Mutations Strengthened SARS-CoV-2 Infectivity --"... It is interesting to note that overall, infectivity-strengthening mutations grow faster than infectivity-weakening mutations, which also reveals that SARS-CoV-2 subtypes having infectivity-strengthening mutations are able to infect more people. Specifically, frequencies of S477N, N439K, V483A, and V367F are higher than those of other mutations, indicating these mutations have a stronger transmission capacity..."
"...Through a normal mode analysis (NMA) we identified the highly flexible region in the receptor binding domain (RBD) of SARS-CoV-2, starting from residue 475 up to residue 485. Structurally, the position S477 shows the highest flexibility among them. At the same time, S477 is hitherto the most frequently exchanged amino acid residue in the RBDs of SARS-CoV-2 mutants. Therefore, using MD simulations, we have investigated the role of S477 and its two frequent mutations (S477G and S477N) at the RBD during the binding to hACE2. We found that the amino acid exchanges S477G and S477N strengthen the binding of the SARS-COV-2 spike with the hACE2 receptor...."
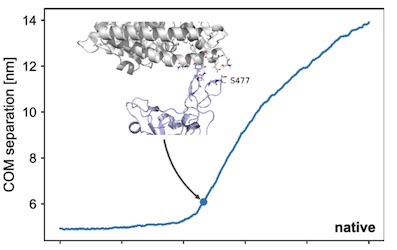
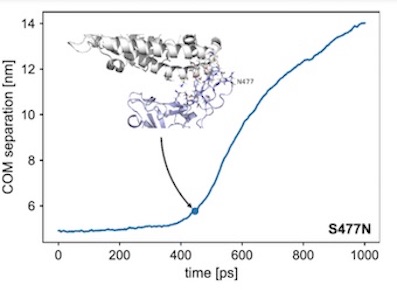
Ball-and-stick representation. Structures and values for the COM (Center of Mass) separation were extracted from an SMD simulation with a spring force constant of 250 kJ/mol/nm2. The breakage point was defined as a COM separation of 1 nm larger than the separation at the start of the simulation.
Sars-CoV-2 Variants
The novel coronavirus SARS-CoV2
B.1.1.7 and B.1.525 UK variant
B.1.427/B.1.429, California QP77P Mutation
B.1.526, NY E484K or S477N Mutations
Molecules of Disease
-
Small Molecules
Cholesterol
Nicotine
Trans Fatty Acid
Alcohol
Acetyaldehyde
- Proteins
Cytokines
Hemoglobin S
Prions
Botulinum Toxin
Explain it with Molecules
- Why is water such a good solvent?
- Why does ice float?
- Why do solids, liquids and gases behave differently?
- What is the geometry of methane?
- What's the difference between alpha and beta glucose?
- How does caffeine work in the brain?
- How does soap work?
- What is the difference between sucrose and fructose?
- Why is carbon monoxide so dangerous?
- Why is graphite so soft if it is made of only carbon?
- What is the difference between Carbyne and Graphite?
- Why is the fullerene and similar structures the cornerstone of nanotechnology?
- How big is a nanotube?
- Why does table salt have a cubic crystal shape?
- What is the structure of the benzene molecule?
- Why do carcinogens cause cancer?
- What causes Sickle Cell Anemia?
- What is the difference between sodium nitrite and nitrate?
- How do drugs work?